V. LLM-in-the-Loop CQL execution with unstructured data and FHIR terminology support
Here is how you can run the CQL execution with unstructured data and FHIR terminology support using our LLM-in-the-Loop stack as presented at AMIA #CIC25.
Bringing Digital health & Gen AI research to life!
Here is how you can run the CQL execution with unstructured data and FHIR terminology support using our LLM-in-the-Loop stack as presented at AMIA #CIC25.
The GitHub repository below is a fork of the CQL Execution Framework, which provides a TypeScript/JavaScript library for executing Clinical Quality Language (CQL) artifacts expressed as JSON ELM. The fork introduces an experimental feature supporting LLM-based assertion checking on DocumentReference. The framework enables execution of CQL logic within different data models, such as QDM and FHIR, but does not provide direct support for data models or terminology services. The library implements various features from CQL 1.4 and 1.5 but has some limitations, such as incomplete support for specific datatypes and functions.
TL; DR: From my personal experiments (on an 8-year-old, i5 laptop with 16 GB RAM), locally hosted LLMs are extremely useful for many tasks that do not require much model-captured knowledge.

Image credit: I, Luc Viatour, CC BY-SA 3.0 https://creativecommons.org/licenses/by-sa/3.0, via Wikimedia Commons
The era of relying solely on large language models (LLMs) for all-encompassing knowledge is evolving. As technology advances, the focus shifts towards more specialized and integrated systems. These systems combine the strengths of LLMs with real-time data access, domain-specific expertise, and interactive capabilities. This evolution aims to provide more accurate, context-aware, and up-to-date information, saving us time and addressing the limitations of static model knowledge.
I have started to realize that LLMs are more useful as language assistants who can summarize documents, write discharge summaries, and find relevant information from a patient’s medical record. The last one still has several unsolved limitations, and reliable diagnostic (or other) decision-making is still in the (distant?) future. In short, LLMs are becoming increasingly useful in healthcare as time-saving tools, but they are unlikely to replace us doctors as decision-makers soon. That raises an important question; Do locally hosted LLMs (or even the smaller models) have a role to play? I believe they do!
Locally hosted large language models (LLMs) offer several key benefits. First, they provide enhanced data privacy and security, as all data remains on your local infrastructure, reducing the risk of breaches and unauthorized access. Second, they allow for greater customization and control over the hardware, software, and data used, enabling more tailored solutions. Additionally, locally hosted LLMs can operate offline, making them valuable in areas with unreliable internet access. Finally, they can reduce latency and potentially lower costs if you already have the necessary hardware. These advantages make locally hosted LLMs an attractive option for many users.
The accessibility and ease of use offered by modern, user-friendly platforms like OLLAMA are significantly lowering the barriers for individuals seeking technical expertise in self-hosting large language models (LLMs). The availability of a range of open-source models on Hugging Face lowers the barrier even further.
I have been doing some personal experiments with Ollama (on docker), Microsoft’s phi3: mini (language model) and all-minilm (embedding model), and I must say I am pleasantly surprised by the results! I have been using an 8-year-old, i5 laptop with 16 GB RAM. I have been using it as part of a project for democratizing Gen AI in healthcare, especially for resource-deprived areas (more about it here), and it does a decent job of vectorizing health records and answering questions based on RAG. I also made a helpful personal writing assistant that is RAG-based. I am curious to know if anybody else in my network is doing similar experiments with locally hosted LLMs on personal hardware.
LangChain is a free and accessible coordination framework for building applications that rely on large language models (LLMs). Although it is widely used, it sometimes receives critiques such as being complex, insecure, unscalable, and hard to maintain. As a novel framework, some of these critiques might be valid, but they might also be a strategy by the dominant LLM actors to regain power from the rebels.
The well-known machine learning frameworks PyTorch and Tensorflow are from the major players who also own some of the largest and most powerful LLMs in the market. By offering these frameworks for free, they can attract more developers and researchers to use their LLMs and platforms and gain more data and insights from them. They can also shape the standards and norms of the LLM ecosystem and influence the direction of future research and innovation.
It may not be the case that the major actors are actively trying to discredit LangChain, but some trends are worth noting. A common misconception is that LLM’s shortcomings are due to LangChain. You would often hear about LangChain hallucinating! Another frequent strategy is to confuse the discussion by bringing conflicting terms to the more widely used LangChain vocabulary. SDKs from major actors (deliberately) attempt to substitute their own syntaxes for LangChain’s.
You might be bewildered after a conference run by the main players, and that could be part of their plan to make you dependent on their products. My approach is to use a mind map to keep track of the LLM landscape and refer to that when suggesting LLM solutions. It also helps to have a list of open-source implementations of common patterns.
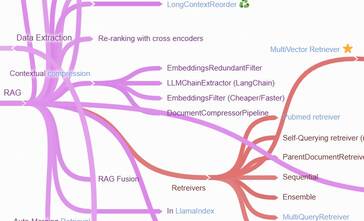
I have also noticed that the big players are gradually giving up and embarrassing the LangChain paradigms. I feel that despite LangChain’s limitations, it is here to stay! What do you think?
TLDR: The targeted distillation method described may be useful for creating an LLM-based cTakes alternative for Named Entity Recognition. However, the recipe is not available yet.

Image credit: Wikimedia
Named Entity Recognition is essential in clinical documents because it enhances patient safety, supports efficient healthcare workflows, aids in research and analytics, and ensures compliance with regulations. It enables healthcare organizations to harness the valuable information contained in clinical documents for improved patient care and outcomes.
Though Large Language Models (LLMs) can perform Named Entity Recognition (NER), the capability can be improved by fine-tuning, where you provide the model with input text that contains named entities and their associated labels. The model learns to recognize these entities and classify them into predefined categories. However, as described before fine-tuning Large Language Models (LLMs) is challenging due to the need for substantial, high-quality labelled data, the risk of overfitting on limited datasets, complex hyperparameter tuning, the requirement for computational resources, domain adaptation difficulties, ethical considerations, the interpretability of results, and the necessity of defining appropriate evaluation metrics.
Targeted distillation of Large Language Models (LLMs) is a process where a smaller model is trained to mimic the behaviour of a larger, pre-trained LLM but only for specific tasks or domains. It distills the essential knowledge of the LLM, making it more efficient and suitable for particular applications, reducing computational demands.
This paper described targeted distillation with mission-focused instruction tuning to train student models that can excel in a broad application class. The authors present a general recipe for such targeted distillation from LLMs and demonstrate that for open-domain NER. Their recipe may be useful for creating efficient distilled models that can perform NER on clinical documents, a potential alternative to cTakes. Though the authors have open-sourced their generic UniversalNER model, they haven’t released the distillation recipe code yet.
REF: Zhou, W., Zhang, S., Gu, Y., Chen, M., & Poon, H. (2023). UniversalNER: Targeted Distillation from Large Language Models for Open Named Entity Recognition. ArXiv. /abs/2308.03279
Deploying large language models (LLMs) can be difficult because they require a lot of memory and computing power to run efficiently. Companies want to create smaller task-specific LLMs that are cheap and easy to deploy. Such small models may even be more interpretable, an important consideration in healthcare.

Distilling LLMs refers to the process of training a smaller, more efficient model to mimic the behaviour of a larger, more complex LLM. This is done by training the smaller model on the same task as the larger model but using the predictions of the larger model as “soft targets” or guidance during training. The goal of distillation is to transfer the knowledge and capabilities of the larger model to the smaller model, without requiring the same level of computational resources.
Distilling step-by-step is an efficient distillation method proposed by Google that requires less amount of training data. The intuition is that the use of rationale generated by a chain of thought prompting along with labels during training, thereby framing it as multi-task learning, improves distillation performance. We can use ground truth labels or use a teacher LLM to generate the labels and rationale. Ground truth labels are the correct labels for the data, and they are typically obtained from human annotators. The rationale for each label can be generated by using the model to generate a short explanation for why the model predicted that label.
The paper on the method is here and the repository is here. I have converted the code from the original repository into a tool that can be used to distill any seq2seq model into a smaller model based on a generic schema. See the repository below. The original paper uses Google’s T5-v1 model, which is a large-scale language model that was developed by Google. It is part of the T5 (Text-to-Text Transfer Transformer) family of models and is based on the Transformer architecture. You can find more open-source base models for distilling on huggingface. The next plan is to use this method to create a model that can predict the FHIR filter for this repository.
I will update this post regularly with my findings and notes on distilling models. Also, please check out my post on NLP tools in healthcare.
OpenFaaS is an open-source framework for building serverless functions with containers. Serverless functions are pieces of code that are executed in response to a specific event, such as an HTTP request or a message being added to a queue. These functions are typically short-lived and only run when needed, which makes them a cost-effective and scalable way to build cloud-native applications. OpenFaaS makes it easy to build, deploy, and manage serverless functions. OpenFaaS CLI minimizes the need to write boilerplate code. You can write code in any supported language and deploy it to any cloud provider. It provides a set of base containers that encapsulates the ‘function’ with a webserver that exposes its HTTP service on port 8080 (incidentally the default port for Google Cloud Run). OpenFaaS containers can be directly deployed on Google Cloud Run and with the faas CLI on any cloud provider.
Kubeflow is a toolkit for building and deploying machine learning models on Kubernetes. Kubeflow is designed to make it easy to build, deploy, and manage end-to-end machine learning pipelines, from data preparation and model training to serving predictions and implementing custom logic. It can be used with any machine learning framework or library. Google’s Vertex AI platform can run Kubeflow pipelines. Kubeflow pipeline components are self-contained code that can perform a step in the machine learning workflow. They are packaged as a docker container and pushed to a container registry that the Kubernetes cluster can access. A Kubeflow component is a containerized command line application that takes input and output as command line arguments.
OpenFaaS containers expose HTTP services, while Kubeflow containers provide CLI services. That introduces the possibility of tweaking OpenFaaS containers to support CLI invocation, making the same containers usable as Kubeflow components. Below I explain how a minor tweak in the OpenFaaS templates can enable this.
Let me take the OpenFaaS golang template as an example. The same principle applies to other language templates as well. In the golang-middleware’s main.go, the following lines set the main route and start the server. This exposes the function as a service when the container is deployed on Cloud Run.
http.HandleFunc("/", function.Handle),
listenUntilShutdown(s, healthInterval, writeTimeout) I have added the following lines [see on GitHub] that expose the same function on the command line for Kubeflow.
if len(os.Args) < 2 {,
listenUntilShutdown(s, healthInterval, writeTimeout)
} else {
dat, _ := os.ReadFile(os.Args[1])
_dat := function.HandleFile(dat)
_ = os.WriteFile(os.Args[2], _dat, 0777)
} If the input and output file names are supplied on the command line as in kubeflow, it reads from and writes to those files. The kubeflow component definition is as below:
implementation:
container:
image: <image:version>
command: [
'sh',
'-c',
'mkdir --parents $(dirname "$1") && /home/app/handler "$0" "$1"',
]
args: [{inputPath: Input 1}, {outputPath: Output 1}]With this simple tweak, we can use the same container to host the function on any cloud provider as serverless functions and Kubeflow components. You can pull the modified template from the repo below.
golang-http-template (this link opens in a new window) by dermatologist (this link opens in a new window)
Golang templates for OpenFaaS using HTTP extensions
This post is meant to be an instruction guide for healthcare professionals who would like to join my projects on GitHub.

Contribution is not always coding. You can clean up stuff, add documentation, instructions for others to follow etc. Issues and feature requests should be posted under the ‘issues’ tab and general discussions under the ‘Discussions’ tab if one is available.
There’s no better time than now to choose a repo to contribute!
The need for computerized clinical decision support is becoming increasingly obvious with the COVID-19 pandemic. The initial emphasis has been on ‘replacing’ the clinician which for a variety of reasons is impossible or impractical. Pragmatically, clinical decision support systems could provide clinical knowledge support for clinicians to make time-sensitive decisions with whatever information they have at the point of patient care.
Providing clinical decision support requires some formal way of representing clinical knowledge and complex algorithms for sophisticated inference. In knowledge management terms, the information requires to be transformed into actionable knowledge. Knowledge needs to be represented and stored in a way conducive to easy inference (knowledge reuse)1. I have been exploring this domain for a considerable period of time, from ontologies to RDF datasets. With the advent of popular graph databases (especially Neo4J ), this seems to be a good knowledge representation method for clinical purposes.
To cut a long story short, I have been working on building a suite of JAVA libraries to support knowledge extraction, annotation and transformation to a graph schema for inference. I have not open-source it yet as I have not decided on what license to use. However, I am posting some preliminary information here to assess interest. Please give me a shout, if you share an interest or see some potential applications for this. As always, I am open to collaboration.
The JAVA package consists of three modules. The ‘library’ module wraps the NCBI’s E-Utils API to harvest published article abstracts if that is your knowledge source. Though data extraction from the clinical notes in EMR’s is a recent trend, it is challenging because of unstructured data and lack of interoperability. The ‘qtakes’ module provides a programmable interface to my quick-ctakes or the quarkus based apache ctakes, a fast clinical text annotation engine. Finally, the graph module provides the Neo4J models, repositories and services for abstracting as a knowledge graph.
The clinical knowledge graph (ckb) consists of entities such as Disease, Treatment and Anatomy and appropriate relationships and cypher queries are defined. The module exposes services that can be consumed by JAVA applications. It will be available as a maven artifact once I complete it.
UPDATE: May 30, 2021: The library (ckblib) is now available under MPL 2.0 license (see below). Feel free to use it in your research.
First posted on CanEHealth.com
The provincial government is building a connected health care system centred around patients, families and caregivers through the newly established OHTs. As disparate healthcare and public health teams move towards a unified structure, there is a growing need to reconsider our information system strategy. Most off the shelf solutions are pricey, while open-source solutions such as DHIS2 is not popular in Canada. Some of the public health units have existing systems, and it will be too resource-intensive to switch to another system. The interoperability challenge needs an innovative solution, beyond finding the single, provincial EMR.

We have written about the theoretical aspects, especially the need to envision public health information systems separate from an EMR. In this working paper, we propose a maturity model for PHIS and offer some pragmatic recommendations for dealing with the common challenges faced by public health teams.
Below is a demo project on GitHub from the data-intel lab that showcases a potential solution for a scalable data warehouse for health information system integration. Public health databases are vital for the community for efficient planning, surveillance and effective interventions. Public health data needs to be integrated at various levels for effective policymaking. PHIS-DW adopts FHIR as the data model for storage with the integrated Elasticsearch stack. Kibana provides the visualization engine. PHIS-DW can support complex algorithms for disease surveillance such as machine learning methods, hidden Markov models, and Bayesian to multivariate analytics. PHIS-DW is work in progress and code contributions are welcome. We intend to use Bunsen to integrate PHIS-DW with Apache Spark for big data applications.
FHIR has some advantages as a data persistence schema for public health. Apart from its popularity, the FHIR bundle makes it possible to send observations to FHIR servers without the associated patient resource, thereby ensuring reasonable privacy. This is especially useful in the surveillance of pandemics such as COVID19. Some useful yet complicated integrations with OSCAR EMR and DHIS2 is under consideration. If any of the OHTs find our approach interesting, give us a shout.
BTW, have you seen Drishti, our framework for FHIR based behavioural intervention?